Pyro Sequencing
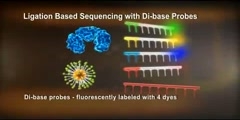
Pyrosequencing™ is sequencing by synthesis, a simple to use technique for accurate and quantitative analysis of DNA sequences.Pyrosequencing is a method of DNA sequencing (determining the order of nucleotides in DNA) based on the "sequencing by synthesis" principle, which relies on detection of pyrophosphate release on nucleotide incorporation rather than chain termination with dideoxynucleotides.The technique was developed by Pål Nyrén and his student Mostafa Ronaghi at the Royal Institute of Technology in Stockholm in 1996.Procedure "Sequencing by synthesis" involves taking a single strand of the DNA to be sequenced and then synthesizing its complementary strand enzymatically. The Pyrosequencing method is based on detecting the activity of DNA polymerase (a DNA synthesizing enzyme) with another chemiluminescent enzyme. Essentially, the method allows sequencing of a single strand of DNA by synthesizing the complementary strand along it, one base pair at a time, and detecting which base was actually added at each step. The template DNA is immobilized, and solutions of A, C, G, and T nucleotides are added and removed after the reaction, sequentially. Light is produced only when the nucleotide solution complements the first unpaired base of the template. The sequence of solutions which produce chemiluminescent signals allows the determination of the sequence of the template. ssDNA template is hybridized to a sequencing primer and incubated with the enzymes DNA polymerase, ATP sulfurylase, luciferase and apyrase, and with the substrates adenosine 5´ phosphosulfate (APS) and luciferin. 1. The addition of one of the four deoxynucleotide triphosphates (dNTPs)(in the case of dATP we add dATPαS which is not a substrate for a luciferase) initiates the second step. DNA polymerase incorporates the correct, complementary dNTPs onto the template. This incorporation releases pyrophosphate (PPi) stoichiometrically. 2. ATP sulfurylase quantitatively converts PPi to ATP in the presence of adenosine 5´ phosphosulfate. This ATP acts as fuel to the luciferase-mediated conversion of luciferin to oxyluciferin that generates visible light in amounts that are proportional to the amount of ATP. The light produced in the luciferase-catalyzed reaction is detected by a camera and analyzed in a program. 3. Unincorporated nucleotides and ATP are degraded by the apyrase, and the reaction can restart with another nucleotide. Currently, a limitation of the method is that the lengths of individual reads of DNA sequence are in the neighborhood of 300-500 nucleotides, shorter than the 800-1000 obtainable with chain termination methods (e.g. Sanger sequencing). This can make the process of genome assembly more difficult, particularly for sequence containing a large amount of repetitive DNA. As of 2007, pyrosequencing is most commonly used for resequencing or sequencing of genomes for which the sequence of a close relative is already available. The templates for pyrosequencing can be made both by solid phase template preparation (Streptavidin coated magnetic beads) and enzymatic template preparation (Apyrase+Exonuclease).
Channels: Genetics Molecular Genetics
Tags: sinhhocvietnam.com vnbio.com
Uploaded by: sikadi ( Send Message ) on 07-07-2009.
Duration: 1m 53s