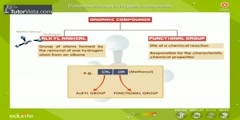
Proteomics Functional Genomics at DNA and RNA
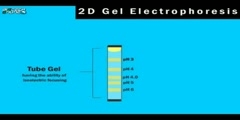
This video describes about proteomics functional genomics at DNA RNA level transcrip. Functional Genomics At DNA And RNA Level: We know that for a large number of organisms a complete genome sequence is now available, so in functional genomics by the help of genomic sequence we can excess every single gene to study and evaluate its function at systemic level Transcriptomics: Functional genomics sequences that are based on DNA sequencing and are used to study mRNA expression profiles on a biological scale are known as transcriptomics. The expression profile of a gene can reveal a lot about its functional role in the cell and can also help to identify functional links to other genes. For example, the expression of many genes is restricted to specific cells or developing structures, often showing that these genes have particular functions in those places other genes are expressed in response to external stimuli. Furthermore mutating one gene may affect the expression profile of the others, helping to link those genes into functional pathways and networks. The two major technologies for the large scale expression analysis that emerged from genomics were large scale complementary cDNA sequencing sampling based on standard DNA sequencing methods and the use of DNA array for expression analysis by hybridization. Sequence sampling: Sequence sampling is the direct and fastest way to study transcriptome. In one of the basic approach clones are picked up from cDNA library then theses clones are identified by their comparison with 200 to 300 bp sequence obtained from sequence databases. By the help of this comparison abundance of each clone will represent the abundance of corresponding transcript in the transcriptome of original biological material now if enough gene are cloned then by statiscal analysis comparison can be made across two or more samples if suitable cDNA library is available. This approach has been used to identify differentially expressed genes but it is expensive because large scale sequencing is required. Several techniques have been developed for high throughput signature recognition (very short sequence samples) but the one that has the most impact thus far is Serial Analysis of Gene Expression (SAGE) SAGE: SAGE is a technique that allow a rapid detailed analysis of thousands of transcripts in a cell. SAGE Procedure: Firstly, polyadenylated RNA is extracted from the cell by the help of enzyme reverse transcriptase we make a DNA strand that is complimentary to RNA. This DNA is then converted to double stranded DNA molecule by the help of streptavidin beads. Once cDNA has been created it is cleaved by using as anchoring enzyme (i.e. used to cut specific 4bp DNA sequence). The cut cDNA is then bound to streptavidin beads by virtue of a multiple thymidine (T's) at its 3' end thereby immobilizing it. The sample of bound cDNA is divided in half and ligated to linker A & B to create ditags. These ditags have linker A on one end and linker B on the other end, and both transcript tags are adjacent to one another in the middle, these ditags are then amplified by PCR. Once amplified they are then cleaved using anchoring enzymes again, this has two effects first is that it release the linkers from either end, and secondly it creates sticky ends in this way all the ditags generated are linked to produce one long string of tags. The collection of tags is introduced into a vector to be cloned and sequenced. DNA Microarray: The method of choice in transcriptomics is the use of DNA micro array. Procedure: In this process firstly we isolate the messenger RNA from two samples. Then mRNA is reverse transcribed into more stable cDNA and for comparative study one of the sample is fluorescently labeled red while the other one is labeled green. At this point in microarray experiment, the two cDNA samples are combined and applied on the microarray chip. The cDNA binds with their complementary sequences on the chip. After hybridization the chip is put into a laser scanner where laser activate the labeled cDNA samples. A computer captures this information and calculates the ratio of red and green on each spot which indicates the expression percentage of both genes.
Channels: Genetics
Tags: Proteomics functional genomics at DNA RNA level transcrip
Uploaded by: juliarobertscool ( Send Message ) on 11-10-2012.
Duration: 5m 26s